This is a repository for PyZebrascope, an open-source Python platform for brain-wide neural activity imaging in zebrafish.
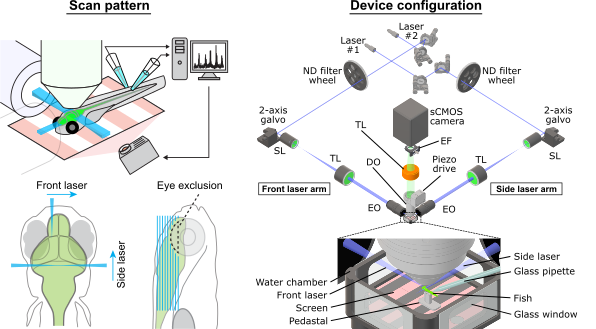
Light-sheet microscopy for whole-brain neural activity imaging in zebrafish requires specific features not available in commercially distributed microscopes. Such features include scanning of two orthogonal excitation beams and eye damage prevention. These particular requirements, as well as numerous device parameters required for the experimenter to manipulate, have been the main bottlenecks for disseminating / developing technology and performing daily experiments. We developed PyZebrascope to address these issues.
PyZebrascope is a high-level interface designed for neural activity imaging experiments based on light-sheet microscopy in zebrafish. It incorporates functionalities from other open-source microscopy projects such as μManager and additional modules for device control, signal processing, computer vision, and machine learning.
A list of hardware that we used to develop PyZebrascope is described in detail in our manuscript https://www.frontiersin.org/articles/10.3389/fcell.2022.875044/full
PyZebrascope requires the preinstallation of the following hardware drivers and low-level interfaces:
- Hardware drivers from manufacturers for miscellaneous devices, including cameras
- DAQmx driver from National Instruments (https://www.ni.com/en-il/support/downloads/drivers/download.ni-daqmx.html)
- μManager (https://micro-manager.org/) for camera interface
- CUDA toolkit (https://developer.nvidia.com/cuda-toolkit) for fast autofocusing based on nVidia GPU
The below Python packages are required to run PyZebrascope:
- Anaconda package (https://anaconda.org/anaconda/python)
- PyQt5 (https://pypi.org/project/PyQt5/)
- pyqtgraph (https://www.pyqtgraph.org/)
- pymmcore (https://github.com/micro-manager/pymmcore, need to match its version with μManager)
- ni-daqmx for Python (https://nidaqmx-python.readthedocs.io/)
- h5py (https://www.h5py.org/)
- CuPy (https://cupy.dev/, need to match its version with CUDA toolkit)
Note that nVidia GPU board, CUDA toolkit and CuPy are only necessary for speeding up the computation time for the autofocusing feature. We still have CPU-based codes in auto_focusing.py
As of December 2023, we confirmed its functionality in Windows10 (x64) with Anaconda 2023.09 which include below Python packages
-
conda 23.11.0
-
Python 3.11.5
-
numpy 1.26.2
-
spyder 5.4.3
-
PyQt5 5.15.10
-
pyqtgraph 0.13.3
-
nidaqmx 0.9.0
-
h5py 3.9.0
-
pyserial 3.5
-
cupy-cuda12x: 12.3.0 (for CUDA 12)
-
micro-manager-2.0.0
-
pymmcore: 10.1.1.70.6 (not the latest but is compatible with an available Hamamatsu μManager API, which requires Device v70 and Module v10)
PyZebrascope supports two laser systems, up to three scanning arms for the front and side illumination of excitation beams, a piezoelectric objective scanner and two sCMOS cameras for multicolor imaging. Below is a configuration with which we tested PyZebrascope.
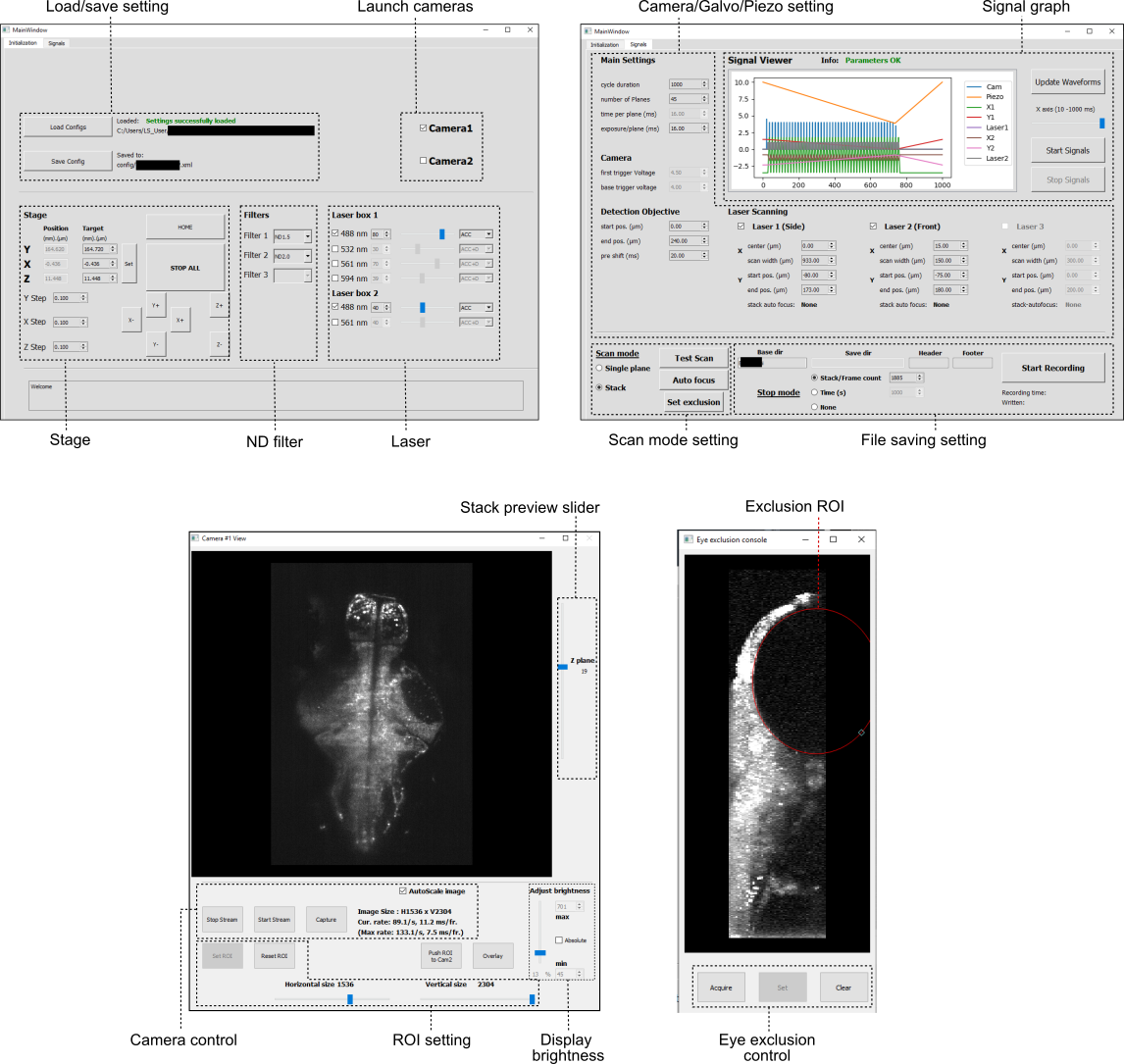
PyZebrascope has two main tabbed interfaces with a camera view window. Additionally, it has an interface to set a laser exclusion area for eye damage prevention.
Image data from cameras are processed like below. Three Qthreads (Reader, Writer, CamView) run in parallel for each camera to support file writing and image previewing. This structure achieves over 800 MB/s writing performances on a fast NVMe drive (Micron 9300) or SSD RAID system while maintaining a stable resource usage of CPU and system memory.
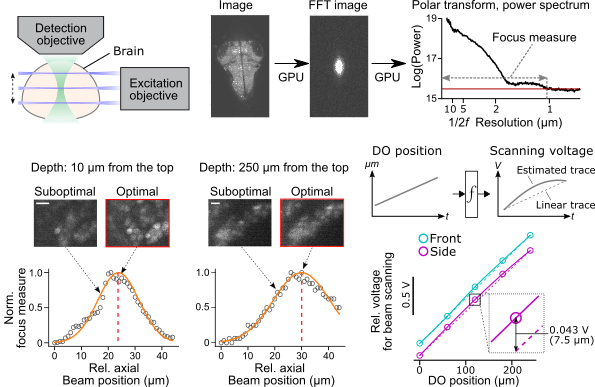
The alignment of the excitation beam to the focus of the detection objective is a time-consuming process for users. We implemented in auto_focusing.py a module for automatically aligning the side laser position to the position of the detection objective lens. It also works for volumetric scans (5-point sampling between the start and the end position) and two excitation arms to fully compensate for the nonlinear relationship between the position of the detection objective positions and analog voltage input to scanning galvanometers.
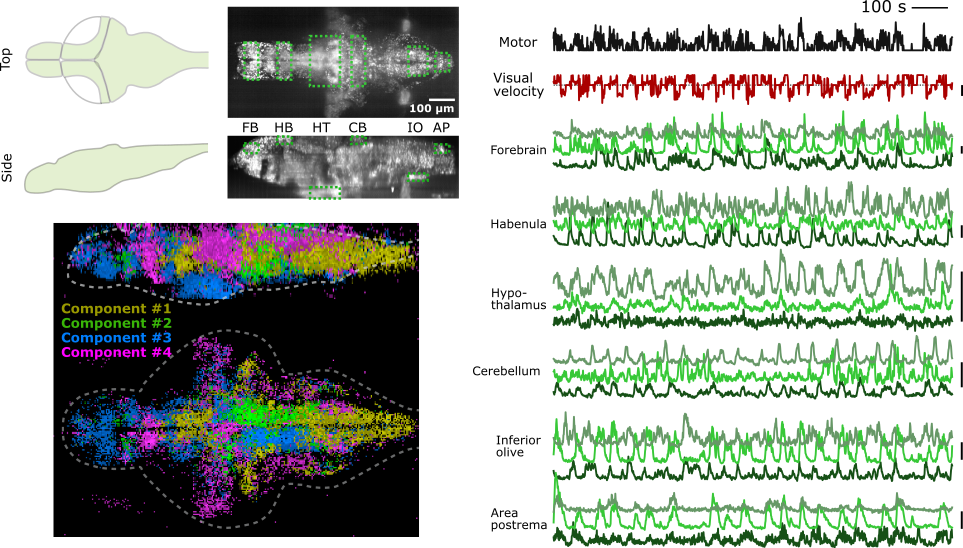
We were able to achieve whole-brain imaging at cellular resolution in a zebrafish performing a motor learning task (kawashima et al., 2016). The imaging volume covers the extremities of the most dorsal part (cerebellum), the most ventral part (hypothalamus), the most rostral part (forebrain), and the most caudal part (hindbrain) at the speed of 1 Hz, 45 planes, 2304 x 1600 pixel resolution.